News and Media
;
How AI can help diagnose cancers that are otherwise hard to spot
OICR-supported researchers are harnessing the power of artificial intelligence to help diagnose cancer quickly and accurately. The difference between cancerous and non-cancerous tissue can be so subtle it’s almost imperceptible. And yet distinguishing one from the other can mean the difference between life and death. Early detection is critical to treating cancer effectively, and it’s […]
Read the story
Apr 22, 2024
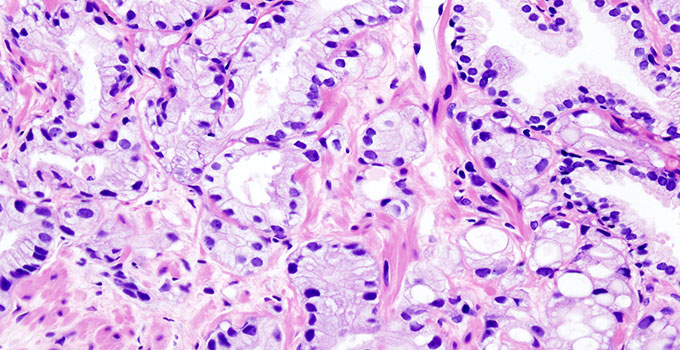
How two cancers make themselves harder to treat by suppressing the immune system
Apr 18, 2024

New “window-of-opportunity” clinical trials explore cutting-edge treatments for cancers of the liver, head and neck
Apr 17, 2024
‘Smart people’ working to make a difference: Co-op student reflects on OICR conference
Apr 16, 2024

Student-developed machine learning model can predict how medicines treat different tumours
Apr 03, 2024

Ontario cancer community comes together at OICR Translational Research Conference
Apr 02, 2024

FDA approves first cell therapy in landmark achievement for personalized medicine
Mar 25, 2024

OICR-supported cancer therapeutics company Fusion Pharmaceuticals acquired by AstraZeneca for $2 billion
Mar 22, 2024

How the Ontario Cancer Research Ethics Board protects participants while streamlining research in the province
Mar 22, 2024

Ask a cancer researcher: Is there a blood test for cancer?
Feb 28, 2024

“Dream team” science helps move Ontario ahead of the curve on hard-to-treat cancers
Feb 27, 2024

Using AI to connect cancer patients with cutting-edge clinical trials
Feb 27, 2024

New PFAC Chair takes lead of growing, evolving community of OICR patient partners
Looking Ahead
Receive the latest news, event invites, funding opportunities and more from the Ontario Institute for Cancer Research.



